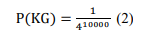IJNSA 04
A SECURE DNA CRYPTOSYSTEM BASED ON STEGANOGRAPHY AND INDEXING CIPHER
Tamer Barakat1 , Nahed Mahmoud1, Ihab. A. Ali2 and Mohamed Hamdi1
1Department of Communications and Electronics, Fayoum University, Egypt
2Department of Communications and Electronics, Helwan University, Egypt
ABSTRACT
One of the essential challenges nowadays; is how to secure data with the increase of its volume as well as its transmission rate. The most frequent approach used to give a high degree of protection, preserve data from hackers, and accomplish multilayer security is steganography combined with encryption. DNA (Deoxyribonucleic Acid) is considered as a new promising carrier for data security while achieving powerful security and maximum protection. In this paper, a secure DNA cryptosystem model which combines steganography with encryption is introduced and categorized into two layers. The original data are hidden in the first layer into a reference DNA based on the insertion method to obtain a fake DNA sequence. In the second layer, this fake DNA sequence, which is the first layer’s output, is encrypted using an indexing cipher to produce an encrypted message in the form of indexes. The proposed model guarantees multilayer security to the secret data with high performance and low-time wasting. It addresses the long-generation key problem of the DNA cryptography. The experimental results assess and validate the theoretical security analysis and model performance.
KEYWORDS
DNA cryptography, Steganography, Indexing cipher, Multilayer security, Reference DNA
1. INTRODUCTION
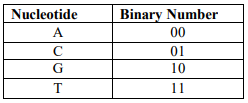
DNA is one of the most recent fields used in cryptography due to the genome carrier’s great storage capacity (1 gm. of DNA can store 106 TB of data) as well as its huge randomness. Random sequences like DNA sequence is considered to be an excellent candidate for use in cryptography [1]. DNA cryptography intends to achieve greater security than conventional cryptography when encrypting data by combining biological and computational properties [2]. DNA is a real carrier of data that can be encoded according to the four-letter alphabet; A, C, G, and T [3]. These bases can be converted into binary according to the binary coding rule mentioned in Table 1.
Table 1. Binary Coding Rule
Information security is an important issue when transferring a secret message between sender and receiver for achieving data confidentiality. Cryptography which can be also referred to as data encryption is used to obtain a secret data writing through enciphering and deciphering using a secret key and a cryptographic algorithm. While the protection of the secret data is the main purpose in security when transmitting data over networks, cryptography isn’t sufficient to provide complete security [4]. Consequently, cryptography must be combined with data hiding techniques or steganography methods to enhance the security and achieve strong data protection[5]. In this paper, the insertion technique is introduced for data hiding method combined with encryption of DNA indexing cipher.
A new direction of DNA computing based on DNA cryptography started with Adleman to introduce a perfect solution for algorithms that requires a huge amount of computations was introduced by [6]. Procedures of DNA OTP encryption schemes are provided as a medium for high computation and for high storage capacity to solve the problem of one-time-pads limited size which used in cryptographic systems, as illustrated in [7]. DNA steganography has been recently found to be a very promising research [8]. In [8], DNA steganography is introduced, in which data is encoded in DNA and hidden in microdots.A pioneer idea which is the essential concept for DNA indexing cipher is presented in [9] to encrypt text and image where The concept to use DNA computing in cryptography is introduced to improve the security of cryptosystems. This is considered the basic idea of our work. The DNA sequence used for encryption is very long; as a result, the average runtime in the experimental results is very high too. DNA chromosomes are used as OTP structure as shown by [10] to encrypt the secret message with the principle of DNA indexing cipher without any experimental results for security analysis and performances. Security level and performances of the algorithm in [10] are analysed by [11] but the secret key is very long and the decryption time was higher than the encryption time. Moreover, the algorithm is unsecured against the related key attack.
In this work, a secure DNA cryptosystem is introduced to eliminate the DNA indexing cipher that appeared in the recent researches by combining another layer of insertion DNA steganography method with it.
2. MOTIVATIONS AND CONTRIBUTIONS
2.1. Motivations
DNA cryptography has significantly become one of the newest technologies in information security. It provides the strength of the genomic database in addition to the basic solutions in cryptography. Therefore, various DNA encryption algorithms are proposed to enhance security in DNA cryptosystem, one of them is the DNA indexing algorithm presented in [11] which ensures a good level of security but, a certain level of vulnerability can be specified to the related-key attack. In the related-key attacks, key transformations can be applied to the secret key. So, the adversary can request plaintext’s encryption using the transformed key until the secret key can be successfully guessed. In besides, the DNA sequence used for encoding and decoding is very long which in turn increases the computation runtime.
2.2. Contributions
To overcome the problem of the DNA indexing algorithm and improve its security, a secure DNA cryptosystem model with double layers of steganography and encryption is presented. In the first layer, the original message will be hidden within reference DNA using the insertion technique to create a fake DNA message. The fake DNA message will next be encrypted using the DNA indexing cipher to generate random indices. The proposed model will entirely secure the original message even if an attacker obtains the decryption key since the original plaintext is still protected and unknown because it is hidden in fake DNA in the first layer.
Therefore, this model provides a multilayer of security against cryptanalytic attacks and as a result, solves the problem of related-key attack by protecting the original message. In addition, the key sequence used in the proposed model is one-third the length that used in [11] which achieves higher performance and evaluation levels for this model.
3. PROPOSED MODEL
Double layer DNA indexing encryption model (DLDI) uses one-time pad symmetric encryption which is considered the essential idea for this model to encrypt each word with one pad for one time. In DLDI model, (OTP) secret key is a genomic DNA sequence downloaded from one of the genomic databases, for example, GenBank, DDBJ, etc. [12]. Encryption is ensured with a genomic key composed of thousands of nucleotide bases. Each genomic key has an ID number which is a unique number with 6 to 8 characters. Recipient must know the ID number which is the key for decryption. In a symmetric key cryptosystem, the encryption key is the same for decryption. So, DLDI model shall protect the original message for an additional time in the case of knowing this key. This is done using the insertion data hiding method which is layer one in DLDI model. In layer two, the encryption process is based upon the indexing encryption presented in [11]. The two layers will be explained below.
3.1. Encryption Process
3.1.1. Layer One
Layer 1 is a modification to the algorithm introduced in [11]. Using the insertion data hiding method in this layer as described in [13] as follows:
Step1: the original message is transformed into decimal ASCII codes then into binary to be the first input to the insertion method.
Step 2: DNA sequence is represented in binary form (S) by using the binary coding which is explained in (Table 1) to be the second input to the insertion method.
Step 3: A number sequence (𝒓𝟏, 𝒓𝟐, … 𝒓𝒕−𝟏) is generated using seed R, the smallest integer t is calculated such that ∑ ri t i=1 ˃ [M] where (t) is the length’s number of the residual part of the message, and a number sequence (𝒌𝟏, 𝒌𝟐, … 𝒌𝒕−𝟏) using seed K is generated too.
Step 4: Message (M) is divided into segments (𝐦𝟏,𝐦𝟐,… 𝒎𝒕−𝟏) with lengths (𝐫𝟏, 𝐫𝟐, … 𝐫𝐭−𝟏) and the residual part will be mt .
Step 5: binary sequence (S) is divided into segments (𝐬𝟏, 𝐬𝟐, … 𝐬𝐭−𝟏) with lengths (𝒌𝟏, 𝒌𝟐, … 𝒌𝒕−𝟏) and neglect the residual part of the sequence.
Step 6: Insert each segment of [M] as a prefix into each segment of the binary sequence (S) then add the segment 𝐦𝐭 at the end of 𝒔𝒕−𝟏.
Step 7: Finally, the inverse function of the binary coding is used to obtain an output with a fake DNA sequence that contains the secret message. Figure 1 explains the DNA insertion method.
Figure 1. DNA Insertion Method
Various algorithms preferred using data hiding after encryption but, in this work, it is preferred to use data hiding before encryption to overcome the problems of DNA indexing encryption while obtaining a high number of substitutions for each plaintext byte without using a very long secret key and achieving a high protection for the secret message.
3.1.2. Layer Two
In this layer, DNA indexing encryption based upon the chromosomal sequence is used as a book to hide the fake DNA output from layer 1. This is done by building a key table that contains all the 256 possible values of any byte. Each byte of those possible values is converted into 4 letter sequence using this principle: 10 01 11 00 → GCTA. Searching one byte at a time through the key sequence containing letters: A, C, G, and T Each time this byte sequence is returned in the chromosomal sequence, its position index is memorized in a vector as one of the possible substitutions of it. All substitution vectors of all bytes are memorized in the key table with size 256xN, where N is variable depending on the number of substitutions of each byte in this table. So, encryption of fake DNA is executed one byte at a time which is a substitution of the byte selected randomly from its retrieved vector in the key table.
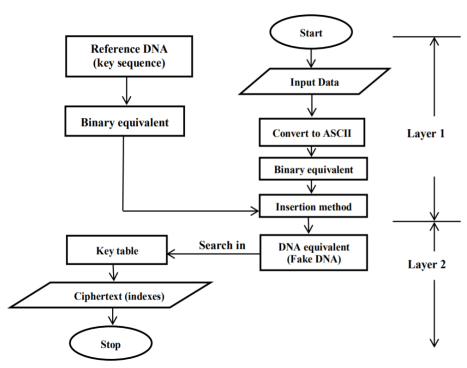
All ciphers that were chosen randomly for all bytes represent the cipher text as presented in Figure 2.
Figure 2. Encryption Process for DLDI Model
In DLDI model there is no need to use a very long chromosomal sequence to obtain a large number of substitutions and choosing randomly one cipher from it for each plain text character like the one used in [11]. Each character from the secret message will be encrypted to more than one cipher from a group of substitutions, explained in the following example.
Using key sequence with length 10,000 bases will be enough to achieve a high level of security and low computational time.
3.1.3. Demonstrative
Example Assume message M: ‘h’, DNA sequence (S): GCGCCCAATACGCAAA ….. with 10,000 bases, message segment length: r=2, and DNA segment length: k=4
- Hiding Layer
Step 1: Convert the plaintext “h” into ASCII: 104 then into binary: 01101000.
Step 2: Get the binary form of (S) using Table 1 to be: 10011001010100001100011001000000……..
Step 3: Divide M into segments with r =2 for each segment: 01, 10, 10, 00.
Step 4: Divide S into segments with k=4 for each segment: 1001, 1001, 0101, 0000, 1100, 0110, 0100, 0000……..till the end of S.
Step 5: Insert each segment from binary M one at a time into the beginning of segments of S till the end of M: 011001 101001 100101 000000.
Step 6: Use the inverse function of binary coding rule to obtain the fake DNA output (S’) CGCGGCGCCAAA
- Encryption Layer
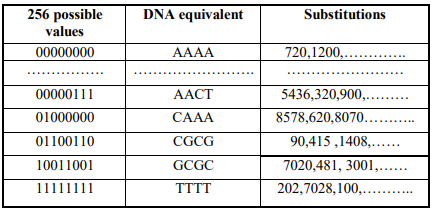
(a) For each byte of this output fake DNA from layer 1, a search in the key table explained in Table 2 is performed to retrieve its vector and choose randomly a substitution to be its cipher. Substitutions of CGCG: 90,415, 1408,…… Ciphertext: 1408 Substitutions of GCGC: 481, 7020 ,3001……… Ciphertext : 3001 Substitutions of CAAA: 8578, 620, 8070,…… Ciphertext: 8578
(b) The ciphertext of “h”: 1408 3001 8578 (more than one cipher for one character from the secret message).
Table 2. Key Table
The maximum number of substitutions that is retrieved in the key table using a DNA sequence with 10,000 bases is 116 substitution values, and the minimum one is 10 substitution values.
3.2. Decryption Process
DLDI model is a symmetric key cryptosystem so; both encryption and decryption use the same key. The receiver has the same coding rule and the values (r and k). Each index from the ciphertext is considered as a pointer to indicate the equivalent DNA byte sequence for every byte from the fake message (S’). Then, it is transformed into binary using the same coding rule and divided into segments with length (r + k) for each one. Finally, extract segments with length r to be the secret message m.
It can be observed that every letter from the original message has more than one substitution from the DNA sequence to be hidden in it and every DNA byte has more than one substitution from the key table which means that the DLDI model guarantees a high level of security for every letter from the original message without using long chromosomal sequence. In addition, ciphertext size is different from plaintext size based upon the DNA segment length where one bit DNA segment length can obtain a fake message before encryption and a different size of ciphertext after it.
4. DATA DESCRIPTION
The quality of our proposed model is evaluated by encrypting different input texts and images with different sizes. These input data are described in Table 3 and Table 4.
Table 3. Dataset for Image
Table 4. Dataset for Text
5. PERFORMANCE EVALUATION AND SECURITY ANALYSIS OF THE MODEL
Two important features that discriminate any cryptographic algorithm from another are its capability to protect data against attacks as well as its speed doing so.
5.1. Performance Evaluation
In this work, Execution time was analysed for three important operations of the DLDI model: key table runtime, encryption and decryption. The key table is executed in 2*256*n operations where 256 is the number of all possible values for each byte and the length of the secret DNA sequence is n. Complexity of the key table computation is O(n), for encryption O(m) and for decryption O(c) where m is the number of plaintext characters and c is the number of ciphertext characters thus the execution time growing rate is linear according to the input size. The experimental results were implemented using MATLAB R2016a and ran on 2.6 GHZ processor under Windows 7 Ultimate.
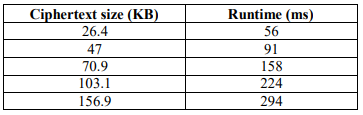
The program was executed for different lengths of n and different sizes of m and c. The computation time measurements are introduced through Table 5 to 9. The execution time of the key table is the same before and after modification. Comparison before and after modification was implemented using a 10,000 bases DNA sequence. Before modification, plaintext size is equal to ciphertext size as explained in Table 6 and Table 7.
Table 5. Computation Runtime of the Key table
Table 6. Encryption Runtime Before Modification
Table 7. Decryption Runtime Before Modification
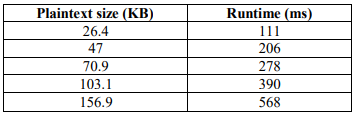
The modification was analysed using the same DNA sequence with message segment length (r=20) and DNA segment length (k=2). The ciphertext size after modification is different for the same Plaintext size used in Table 7.
Table 8. Encryption Runtime after Modification
Table 9. Decryption Runtime after Modification
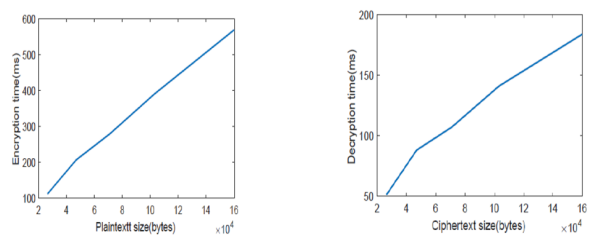
As explained in (Table 9), Decryption Runtime after modification is almost one-half the size before modification especially in large ciphertext size. Consequently, these results proved the high performance of this model. Graphing of the execution time before and after modification is shown below from Figure 3 to Figure 5.
Figure 3. Growing Rate of Key Table execution Time
Figure 4. Growing Rate of Encryption and Decryption Runtime before Modification
Figure 5. Growing rate of Encryption and Decryption Runtime after modification
5.2. Security Analysis for DLDI Model
The security analysis is classified into three main categories: statistical measurements, keyspace (brute force) and cryptanalytic attacks.
5.2.1. Histogram Analysis
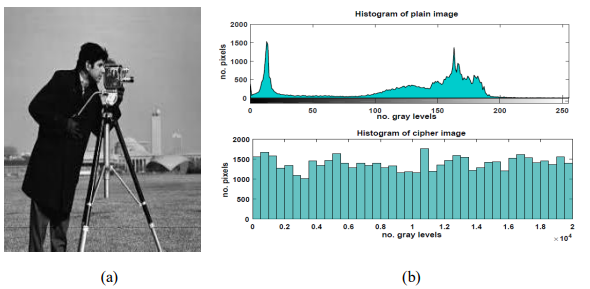
Histogram is one of the statistical measurements that enable visualizing the probability distribution of the signal. With a specified range of values in a signal, the occurrence of each value for it is shown by the histogram [14]. To resist the statistical attacks, a secure encryption system must provide a uniform histogram for the ciphertext [15]. Histograms of plaintext and ciphertext are shown in Figure 6.
Figure 6. Plain image (a) histogram of plain image and encrypted image (b) for DLDI model
Figure 6.b shows that the distribution of the encrypted images is different completely from the distribution of the plain images (Figure 6.a). Hence, our model provides a uniform distribution for the cipher images. In addition, there are no matched patterns between the plaintext and ciphertext histograms thus, a high level of security is ensured here. Histogram analysis was also implemented on a text containing 26400 characters as presented in Figure 7.
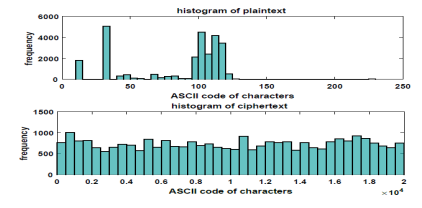
Figure 7. Histogram of a Plaintext and its Ciphertext
It can be noticed that there is a small range of characters in the plaintext while it has a high range of corresponding random cipher in the ciphertext. Hence, it is impossible to recognize the plaintext from the ciphertext.
5.2.2. Entropy Analysis
Entropy is another statistical measurement used to measure the randomness of the ciphertext and the security strength of the signal [16]. Experimental results are presented in Table 10 shows the information entropy analysis before and after modification. Measurements are implemented for text and image files.
Table 10. Plaintext and Ciphertext Entropy analysis before and after modification
Table 10 demonstrates that the entropy of the ciphertext is three times higher than the plaintext entropy for text files and almost twice higher than the plaintext entropy for image. A great value of entropy reaching its maximum value (14 bits/symbol) for the codeword of the ciphertext as, the maximum length of the key sequence is 10000 bases which ensure that it is more effective.
5.2.3. Correlation Analysis
Correlation coefficient (CC) is an important statistical measurement that indicates the degree of dependence between two values like pixels or letters. The range of CC values is from (-1) to (+1) where ±1 refers that the two values are linked to each other (0) indicates that they may be independent. So, CC is desired to be low as possible when analyzing the ciphertext.
Table 11. Plaintext and Ciphertext correlation analysis before and after modification
Table 11 indicates that the CC of the plaintext is 1 or close to 1 while the CC of the ciphertext is close to 0 for DLDI model after modification compared with results in [11] which ensures that the proposed model is effective against different attacks.
5.2.4. Key Space
Key space is one of the main parameters that represent the ability of the algorithm to resist any possible brute-force attack. In DLDI model both encryption and the decryption key is a genetic sequence downloaded from a public database like GenBank which contains millions of DNA sequences reaching around 218642238 sequences [12]. Thus, the key space is equal to the number of all sequences in this large database. Moreover, the genetic sequence used in it is at least 10,000 bases with four letters: A, C, G, and T which means trying 4 10000 possible keys.
5.2.5. Cryptanalytic Attacks
To crack the proposed model, there are two main information must be known: DNA reference used for data hiding layer and the key used for encryption layer.
- The probability of an intruder to have a successful guess about the DNA reference used for data hiding method and DNA binary coding rule as illustrated in [17] but with a change of 2.18 *108 instead of 1.63 *108 for the newest number of DNA sequences in GenBank database is given below:
- The probability of an intruder to have a successful guess about the key used for encryption in this model where there are 4 nucleotides with 10,000 bases for the reference key is:
As a result, the total probability of an intruder to attack the proposed model can be calculated by
combining Equation 1 and Equation 2.
Therefore, combination between data hiding method and DNA indexing cipher makes the proposed model more robust against several attacks.
DLDI model can be resistant to chosen- ciphertext attack as plaintext isn’t ciphered directly but hidden in a fake DNA sequence then ciphered thus makes it very hard to know plaintext from a chosen ciphertext. As it can resist a chosen-ciphertext attack so, it has the capability to resist the other main attacks such that: known-plaintext, chosen-plaintext, and ciphertext-only attack. It guarantees a good level of protection for the original data against related-key attacks as in the case of knowing the encryption key; the adversary can’t know the secret data until the method used for data hiding and number seeds (r, k) are known. Since this model introduces more additional time to protect information.
6. COMPARISON WITH OTHER RECENT WORKS
Various evaluation parameters like entropy, correlation, and key space ensured that the results of our proposed model are better than other encryption algorithms. A comparison of our work with other recent works is shown in Table 12 to Table 14.
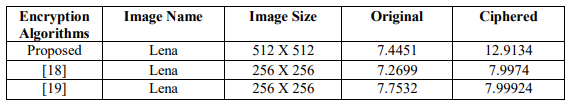
Table 12. Comparison of Information Entropy Analysis Using different Encryption Algorithms
Table 13. Correlation Analysis for Lena Image
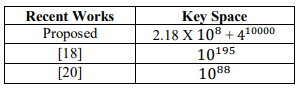
Table 14. Comparison of key Space with other Recent Algorithms
7. CONCLUSION AND FUTURE WORK
In this study, a new DNA-based steganography and encryption is proposed. It enhanced the security strength of the DNA indexing encryption and provided multilayer security to the secret message. Additionally, it eliminated the problem of long key generation which is an essential problem in recent algorithms based on DNA cryptography. It provided maximum protection and perfect hiding to the original data to be secure against related key attacks. Moreover, we achieved a higher time performance. Text and image transmission were implemented and proved with security measurements.
As a future work, we will expand the implementation for video and audio applications. The security proof for related key attack will be introduced as well.
REFERENCES
[1] P. Pavithran, S. Mathew, S. Namasudra, and P. Lorenz, “A novel cryptosystem based on DNA cryptography and randomly generated mealy machine,” Comput. Secur., vol. 104, pp. 102–160, 2020, doi: 10.1016/j.cose.2020.102160.
[2] A. Das, S. K. Sarma, and S. Deka, “Data Security with DNA Cryptography,” in Transactions on Engineering Technologies, 2019, pp. 159–173, doi: 10.1007/978-981-15-8273-8.
[3] M. Mondal and K. S. Ray, “Review on DNA Cryptography,” arXiv Prepr. arXiv1904.05528, pp. 1–31, 2019.
[4] G. Hamed, M. Marey, S. A. El-Sayed, and M. F. Tolba, “Hybrid technique for steganography-based on DNA with n-bits binary coding rule,” in 2015 7th International Conference of Soft Computing and Pattern Recognition (SoCPaR), 2015, pp. 95–102, doi: 10.1109/SOCPAR.2015.7492790.
[5] M. S. Taha, M. S. Mohd Rahim, S. A. Lafta, M. M. Hashim, and H. M. Alzuabidi, “Combination of Steganography and Cryptography: A short Survey,” in IOP Conference Series: Materials Science and Engineering, 2019, vol. 518, no. 5, doi: 10.1088/1757-899X/518/5/052003.
[6] A. Gahlaut, A. Bharti, Y. Dogra, and P. Singh, “DNA based cryptography,” Int. J. Cyber-Security Digit. Forensics, vol. 750, pp. 205–215, 2017, doi: 10.1007/978-981-10-6544-6_20.
[7] J. Zhang and J. Zhang, “DNA based random key generation and management for OTP encryption,” Biosystems, pp. 51–63, 2017.
[8] S. Marwan, A. Shawish, and K. A. Nagaty, “Utilizing DNA Strands for Secured Data-Hiding with High Capacity,” Int. J. Interact. Mob. Technol., vol. 11, no. 2, pp. 88–98, 2017, doi: 10.3991/ijim.v11i2.6565.
[9] S. T. Amin, M. Saeb, and S. El-Gindi, “A DNA-based implementation of yaea encryption algorithm,” in Proceedings of the 2nd IASTED International Conference on Computational Intelligence, 2006, pp. 116–120.
[10] O. Tornea, B. Monica, H. Tatiana, and M.-F. Vaida, “Encryption system with Indexing DNA chromosomes cryptographic algorithm,” IASTED Int. Conf. Biomed. Eng., vol. 680, no. 99, pp. 12– 15, 2010.
[11] O. Tornea and M. E. Borda, “Security and Complexity of a DNA-Based Cipher,” RoEduNet Int. Conf. IEEE, pp. 1–5, 2013, doi: 10.1109/RoEduNet.2013.6511755.
[12] “GenBank and WGS Statistics.” https://www.ncbi.nlm.nih.gov/genbank/statistics/ (accessed Dec. 17, 2020).
[13] H. J. Shiu, K. L. Ng, J. F. Fang, R. C. T. Lee, and C. H. Huang, “Data hiding methods based upon DNA sequences,” Inf. Sci. (Ny)., vol. 180, no. 11, pp. 2196–2208, 2010, [Online]. Available: http://dx.doi.org/10.1016/j.ins.2010.01.030.
[14] D. Balanici, V. Tomsa, M. Borda, and R. Malutan, “Full duplex OTP cryptosystem based on DNA Key for text transmissions.,” Int. Conf. Inf. Technol. Commun. Springer, Cham, pp. 39–48, 2015.
[15] H. Zhu, C. Zhao, X. Zhang, and L. Yang, “An image encryption scheme using generalized Arnold map and affine cipher,” Optik (Stuttg)., vol. 125, no. 22, pp. 6672–6677, 2014, doi: 10.1016/j.ijleo.2014.06.149.
[16] M. Borda, “Fundamentals in information theory and coding,” in Springer Science & Business Media., 2011, pp. 1–504.
[17] S. Marwan, A. Shawish, and K. Nagaty, “An enhanced DNA-based steganography technique with a higher hiding capacity,” in Proceedings of the International Conference on Bioinformatics Models, Methods and Algorithms, 2015, pp. 150–157, doi: 10.5220/0005246501500157.
[18] N. Iqbal, M. Hanif, S. Abbas, M. A. Khan, and Z. Ul Rehman, “Dynamic 3D scrambled image based RGB image encryption scheme using hyperchaotic system and DNA encoding,” J. Inf. Secur. Appl., vol. 58, pp. 1–19, 2021, doi: 10.1016/j.jisa.2021.102809.
[19] K. C. Jithin and S. Sankar, “Colour image encryption algorithm combining, Arnold map, DNA sequence operation, and a Mandelbrot set,” J. Inf. Secur. Appl., vol. 50, pp. 1–22, 2019, doi: 10.1016/j.jisa.2019.102428.
[20] X. Wang, Y. Wang, X. Zhu, and C. Luo, “A novel chaotic algorithm for image encryption utilizing one-time pad based on pixel level and DNA level,” Opt. Lasers Eng., vol. 125, pp. 1–12, 2019, doi: 10.1016/j.optlaseng.2019.105851.